The CaDNAP 13-STR panel
To gain general acceptance forensic DNA testing in animals needs to improve standardization of analysis methods and data interpretation. Recently, the International Society of Forensic Genetics (ISFG) took particular care of this topic by publishing recommendations for forensic non-human DNA analysis following the successful example of human DNA analysis in order to provide a basis for harmonization of the still existing inter-laboratory variability. By following these recommendations we demonstrate the performance of two short tandem repeat (STR) multiplexes for forensic identity testing of canine biological material. Thirteen STRs and two sex-specific markers were selected and validated according to the ISFG guidelines. Population genetic parameters were calculated based on 295 dog samples collected in Austria (124) and Germany (171). A repeat-based nomenclature of the mainly tetrameric STRs and corresponding allelic ladders are presented. All 146 different alleles included in the ladders were sequenced for correct allele calling. Additionally, a canine cell line (DH82-D3167) was evaluated as standard reference material.
Forensic Sci Int Genet 2014 8:90-100
Canine STRs: Allelic ladders
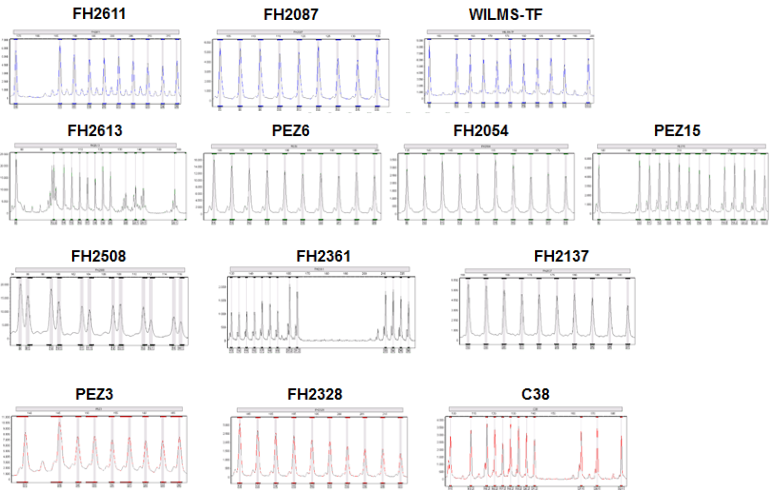
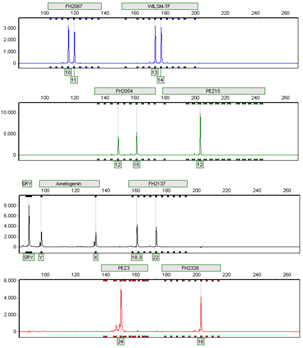
Example of a Canine STR-Multiplex
Breed assignment using the 13 autosomal STR markers
We tested a panel of 13 highly polymorphic canine short tandem repeat (STR) markers for dog breed assignment using 392 dog samples from the 23 most popular breeds in Austria, Germany, and Switzerland. This STR panel had originally been selected for canine identification. The dog breeds sampled in this study featured a population frequency ≥1% and accounted for nearly 57% of the entire pedigree dog population in these three countries. Breed selection was based on a survey comprising records for nearly 1.9 million purebred dogs belonging to more than 500 different breeds. To derive breed membership from STR genotypes, a range of algorithms were used. These methods included discriminant analysis of principal components (DAPC), STRUCTURE, GeneClass2, and the adegenet package for R. STRUCTURE analyses suggested 21 distinct genetic clusters. Differentiation between most breeds was clearly discernable. Fourteen of 23 breeds (61%) exhibited maximum mean cluster membership proportions of more than 0.70 with a highest value of 0.90 found for Cavalier King Charles Spaniels. Dogs of only 6 breeds (26%) failed to consistently show only one major cluster. The DAPC method yielded the best assignment results in all 23 declared breeds with 97.5% assignment success. The frequency-based assignment test also provided a high success rate of 87%. These results indicate the potential viability of dog breed prediction using a well-established and sensitive set of 13 canine STR markers intended for forensic routine use.
Forensic Sci Int Genet 2018 37:126-134
Example of dog breed assignment using GeneClass2
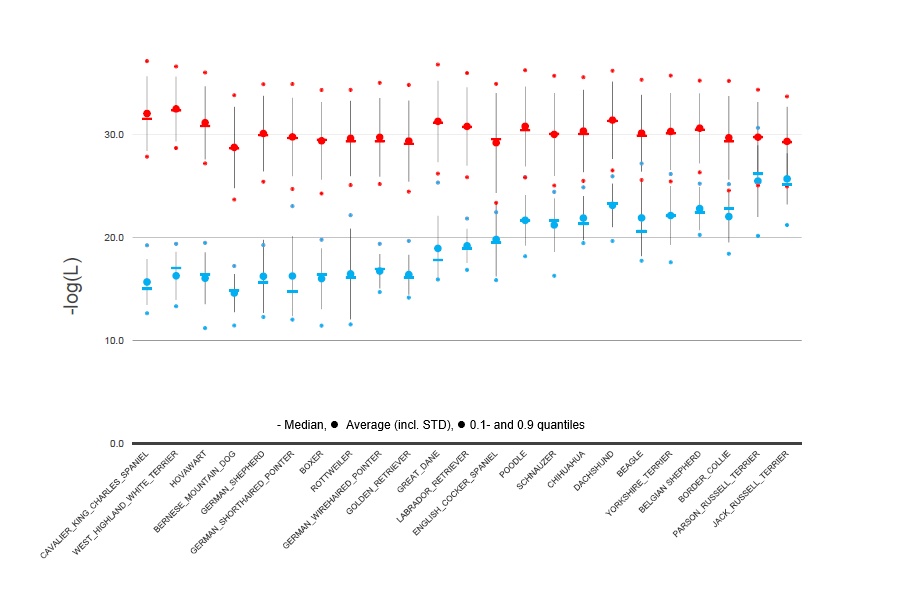
Genotype frequencies in the 23 tested dog breeds: The genotype frequency was calculated with the GeneClass2 software and is shown as the –log (L) for each breed by using the allele frequencies of either the declared breed (open symbols) or, respectively, of the remaining 22 breeds combined (black symbols).